sorting — Sorting Components¶
Sorting components for use in evolutionary algorithms.
As some important orderings in evolutionary computation are only partial, the Python facilities for sorting are not sufficient. Thus, we provide the classes in this module. Lower objective values are always considered better.
-
class
evoalgos.sorting.NotComparableError(object1, object2)¶ Error indicating that two objects cannot be compared.
Inherits from
ValueError.
-
class
evoalgos.sorting.SortingComponent(sort_key=None)¶ Interface for a sorting component.
-
identify_best_group(population)¶ Identify the group of best objective values.
Parameters: population (list of Individual) – The population to partition. Returns: best_group Return type: list of Individual
-
identify_groups(population)¶ Identify groups of equal objective values.
Parameters: population (list of Individual) – The population to partition. Returns: groups Return type: list of list
-
obtain_sort_keys(population)¶ Return a dictionary containing the sort keys for all individuals.
-
sort(population)¶ Sort a population according to the sort keys.
-
-
class
evoalgos.sorting.LexicographicSorting¶ Lexicographic sorting.
-
static
key(individual)¶ Sort key for lexicographic sorting with special treatment of None.
None is replaced with infinity (the worst possible value).
-
static
-
class
evoalgos.sorting.RankSumSorting(ties='average')¶ Rank-sum sorting.
This sorting component ranks the objective values for each objective and uses the sum of ranks as sorting criterion. This approach is a basic variant of the Borda count and is especially suited for many-objective optimization.
-
__init__(ties='average')¶ Constructor.
Parameters: ties (str, optional) – A string determining the treatment of ties in the rank calculation. Possible values are “average”, “min”, “max”, or “first”.
-
obtain_sort_keys(population)¶ Return a dictionary containing the sort keys for all individuals.
-
-
class
evoalgos.sorting.NonDominatedSorting(dominance_relation=None)¶ Non-dominated sorting according to Pareto-dominance.
-
__init__(dominance_relation=None)¶ Constructor.
Parameters: dominance_relation (callable, optional) – A callable that takes two individuals as arguments and returns True or False. Default is dominates().
-
compute_non_dom_front_2d(population)¶ Return the minimal elements in the special case of two objectives.
Does not ensure stability. Only call directly if you are absolutely sure you have two objectives. Run time is
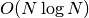 for N
individuals.
For the user it is recommended to call
for N
individuals.
For the user it is recommended to call
identify_best_group()instead.
-
compute_non_dom_front_arbitrary_dim(population)¶ Return the minimal elements for arbitrary dimension.
Does neither ensure stability nor exploit special cases, but can handle an arbitrary number of objectives. Run time is
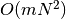 for m objectives and N individuals.
For the user it is recommended to call
for m objectives and N individuals.
For the user it is recommended to call
identify_best_group()instead.
-
static
dominates(individual1, individual2)¶ Evaluate normal Pareto-dominance relation.
None is treated as worst objective value.
-
identify_best_group(population)¶ Return all non-dominated in the given population.
Guarantees stability and exploits 1-D and 2-D special cases if possible.
-
identify_groups(population)¶ Return a partition of the population into non-dominated fronts.
In the identified partition, front i would be non-dominated if fronts 0 to i-1 were not in the population. The worst-case run time is
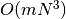 for m objectives and N individuals in the
population. Guarantees stability and exploits 1-D and 2-D special
cases if possible.
for m objectives and N individuals in the
population. Guarantees stability and exploits 1-D and 2-D special
cases if possible.
-
sort(population)¶ Reorder population to be concatenation of non-dominated fronts.
This sorting is stable.
-
static
strongly_dominates(individual1, individual2)¶ Evaluate strong Pareto-dominance relation.
None is treated as worst objective value.
-
static
weakly_dominates(individual1, individual2)¶ Evaluate weak Pareto-dominance relation.
None is treated as worst objective value.
-
-
class
evoalgos.sorting.CrowdingDistanceSorting¶ Sort by non-domination rank and crowding distance.
Implements the sorting order suggested in [Deb2000].
-
sort_front(front, crowding_distances=None)¶ Sort a front according to the crowding distance.
-
-
class
evoalgos.sorting.HyperVolumeContributionSorting(reference_point=None, hv_indicator=None, prefer_boundary_points=True)¶ Sort by non-domination rank and exclusive contribution to the front.
This sorting is required for
HyperVolumeContributionSelection. Further information can be found in [Emmerich2005] and [Naujoks2005].-
__init__(reference_point=None, hv_indicator=None, prefer_boundary_points=True)¶ Constructor.
Parameters: - reference_point (Individual or iterable, optional) – This point is used to bound the hypervolume.
- hv_indicator (HyperVolumeIndicator, optional) – An instance with the interface of
QualityIndicator.FonsecaHyperVolumeis chosen as default. It is only used for three or more objectives. The 2-D special case is treated directly here, to save some time. - prefer_boundary_points (bool, optional) – This flag only pertains to the two-dimensional case. If it is set to True, the two boundary points (but not their potentially existing duplicates) of a 2-D front are put to the front of the sorted population.
-
reference_point¶ The reference point for hypervolume computations.
-
sort_front(front, others=None)¶ Sort a non-dominated front.
Parameters: - front (sequence of Individual) – The individuals to sort.
- others (sequence of Individual, optional) – Other individuals that may influence the exclusive hypervolume contribution of the sorted population, but which are not sorted themselves. Empty by default.
-
sort_front_2d(front, others=None)¶ Sort non-dominated front in the special case of 2 objectives.
Parameters: - front (sequence of Individual) – The individuals to sort.
- others (sequence of Individual, optional) – Other individuals that may influence the exclusive hypervolume contribution of the sorted population, but which are not sorted themselves. Empty by default.
-
-
class
evoalgos.sorting.NearestBetterSorting(dist_matrix_function=None, archive=None, sorting_component=None, use_neighbor_count=False, multiobjective=False)¶ Sort using distances to the nearest better neighbor.
-
__init__(dist_matrix_function=None, archive=None, sorting_component=None, use_neighbor_count=False, multiobjective=False)¶ Constructor.
Parameters: - dist_matrix_function (callable, optional) – A function which takes two iterables of individuals as input
(say, sizes n and m), and returns an
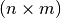 -matrix
of distances as output. Default is Euclidean distance.
-matrix
of distances as output. Default is Euclidean distance. - archive (list of Individual, optional) – Individuals which influence the distance calculations for the sorted population. Empty by default.
- sorting_component (SortingComponent, optional) – This component is used to determine the fitness levels in the
population, by its method identify_groups(). Default is
LexicographicSorting. - use_neighbor_count (bool, optional) – Indicates which criterion should be used for sorting. If False, the sort order is descending by nearest-better neighbor distances. If True, for each individual the neighbors being closer than the nearest-better neighbor are counted. The sort order is then descending by this number. In any case, the order of the sorting component is used as secondary criterion to break ties.
- multiobjective (bool, optional) – Decides if the nearest-better data and objective-related data is combined in a multiobjective fashion (by non-dominated sorting) or by lexicographic sorting.
- dist_matrix_function (callable, optional) – A function which takes two iterables of individuals as input
(say, sizes n and m), and returns an
-
calc_nearest_better_distances(population, others=None, order_dict=None)¶ Calculate the nearest-better distances required for sorting.
Parameters: - population (list of Individual) – The individuals to sort.
- others (list of Individual) – Individuals which influence the distance calculations for the sorted population. Empty by default.
- order_dict (dict, optional) – An empty dictionary into which the fitness levels of each individual are written. So, this additional information can be obtained, if desired.
Returns: min_distances – A dict containing the distance to the nearest-better neighbor for each individual. If no such neighbor exists, the value is infinity.
Return type:
-
count_neighbors_closer_than_nbn(population, others=None, order_dict=None, distances_dict=None)¶ Count the neighbors closer than the nearest better one.
Parameters: - population (list of Individual) – The individuals to sort.
- others (list of Individual) – Individuals which influence the distance calculations for the sorted population. Empty by default.
- order_dict (dict, optional) – An empty dictionary into which the fitness levels of each individual are written, so this additional information can be obtained, if desired.
- distances_dict (dict, optional) – An empty dictionary into which the nearest-better distances are written, so this information can also be obtained through this function call.
Returns: neighbor_counts – A dict containing the number of neighbors closer than the nearest-better one for each individual. If no such neighbor exists, this value is by definition equal to
len(population) + len(others).Return type:
-